Running on your Desktop
Jolecule is also designed to run on your desktop, independent of this website. First you need to download the Jolecule package from Github.com:
jolecule-master.zipYou then need to install Node.js, the Javascript eco-system. Once Node.js is installed, go to the Jolecule directory, and install the modules required:
> npm installThis will include the installation of the Electron package, which enables Desktop mode for Jolecule.
Desktop Electron mode of Jolecule
On the Linux/MacOSx command-line, run from the Jolecule directory:
> ./jol-electron.sh [PDB file] | [directory]
This will run the desktop version of Jolecule that allows you to directly read the PDB files on your drive. As well, there is a useful left-hand panel that will list all PDB structures in the current directory, with the TITLE field from the PDB files.
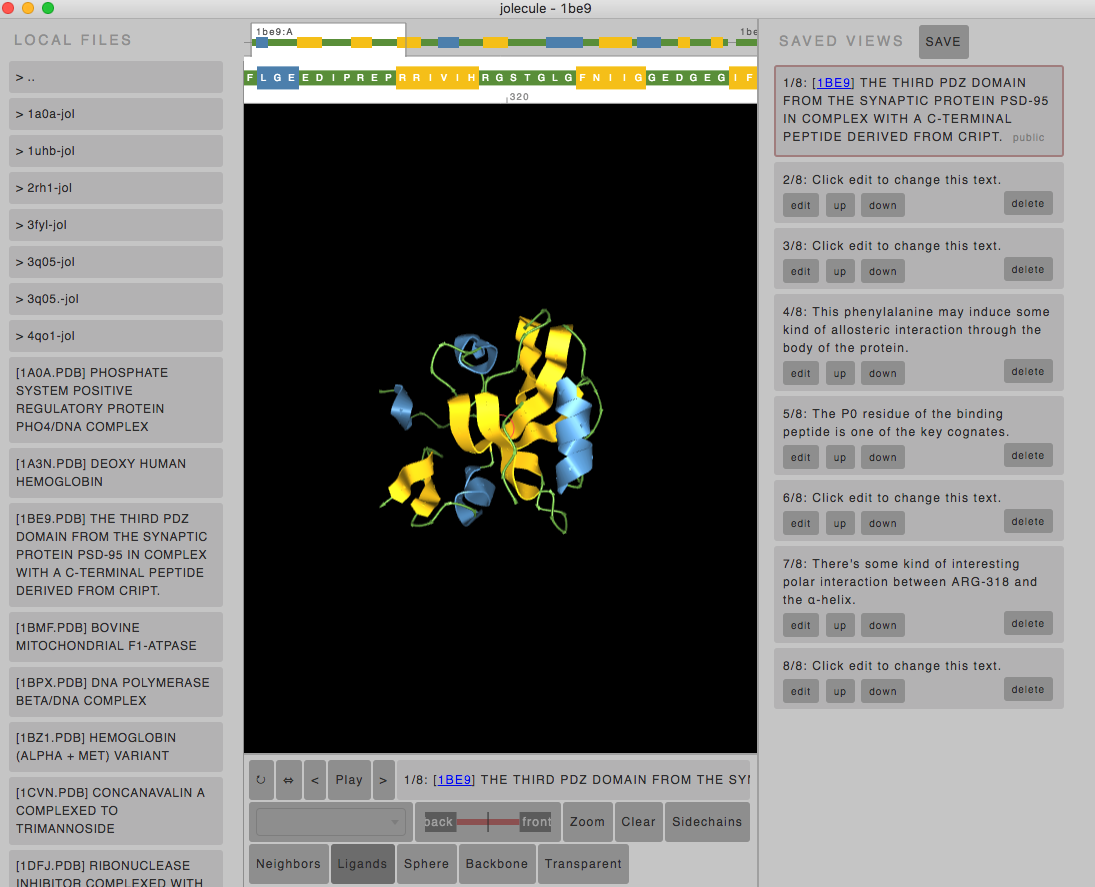
You can easily switch files by clicking on the different PDB entries on the left.
For any PDB file, say 1be9.pdb, any views saved will be written to
the file 1be9.views.json . These views will be automatically
reloaded if the PDB file is later reloaded.
Creating a self-contained Webapp
Once you've created a bunch of views for a PDB structure. You can create a static webapp for the PDB structure and its views. These webapps are self-contained - they can be viewed simply by opening them in a browser. They can be opened locally, remotely on a website, or sent to others.
To create a Jolecule webapp, on the command-line on Linux/MacOSX, in the Jolecule directory, which
contains your PDB file, say 1be9.pdb, and the views file,
say 1be9.views.json, run:
> ./jol-static.js 1be9
This will create a folder 1be9-jol with the webapp, which can be opened in
your browser by opening the following file:
> open 1be9-jol/index.html
There are different ways to setup the webapp, for instance, you can animate a slideshow,
or rotate the protein in a given view. To choose the mode, you can
edit the parameters of the Jolecule widget in
1be9-/jolindex.html
The parameters are described in the embedded widget guide.